Flow Matching for Medical Image Synthesis: Bridging the Gap Between Speed and Quality
MOTFM (Medical Optimal Transport Flow Matching) accelerates medical image synthesis with far fewer sampling steps while maintaining, and often improving, image quality across 2D/3D and class/mask-conditional setups.
Authors: Milad Yazdani, Yasamin Medghalchi, Pooria Ashrafian, Ilker Hacihaliloglu, Dena Shahriari · Affiliation: University of British Columbia, Vancouver, Canada
Abstract (short)
Medical AI often lacks large, high-quality datasets due to privacy and annotation costs. Generative models can help, but diffusion models are slow at inference. MOTFM introduces an Optimal Transport Flow Matching approach that maps source noise to the data distribution along a much straighter path, reducing the number of sampling steps while preserving, and in many cases improving, image quality. The framework supports multiple modalities (e.g., echocardiography and MRI), conditioning mechanisms (class labels and masks), and both 2D and 3D settings.
See arXiv:2503.00266 for the full abstract.
Project news
- May 27, 2025 Accepted to MICCAI 2025.
- Apr 09, 2025 Code released.
- Mar 29, 2025 Paper available on arXiv.
Source: project README.
Method at a glance
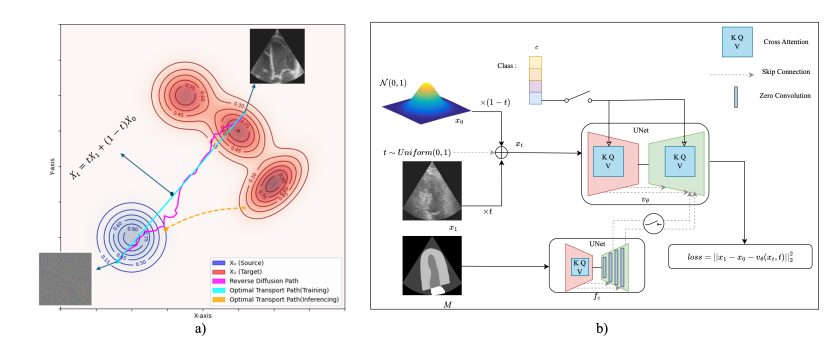
MOTFM trains a UNet-based velocity field with attention to follow an Optimal Transport (OT) path between a simple prior and the target image distribution. Compared to diffusion, the OT path is close to linear, so sampling needs far fewer steps. The design supports: (1) unconditional generation, (2) class-conditional generation via cross-attention, and (3) mask-conditional generation using a second UNet with zero-conv and skip connections; also combinable with class conditioning.
- Architecture: UNet with attention; memory-efficient attention is used to speed training/inference.
- Dimensions: native 2D and 3D variants.
- Conditioning: class labels, masks, or both.
Details: see Sections 2-3 of the paper.
Key results (qualitative & quantitative)
- On echocardiography (CAMUS), MOTFM achieves better distributional and structural metrics than baselines.
- 1-step MOTFM outperforms 10-step DDPM; 10-step MOTFM surpasses 50-step DDPM.
- On 3D MRI (MSD Brain), MOTFM yields stronger 3D-FID/MMD with far fewer steps.
See paper results & tables for exact numbers.
Datasets used
CAMUS (Echo)
2D echocardiography dataset with four- and two-chamber views, widely used for segmentation and generative evaluation.
MSD (Brain MRI)
Medical Segmentation Decathlon, brain tumor task, multi-parametric MRI collection for benchmarking.
Reproducibility
Evaluation follows common distributional and structural metrics; consult the paper for protocols, splits, and exact implementations.
Metrics include FID/3D-FID, KID, CMMD/MMD, IS, (MS-)SSIM.
Quickstart
Install dependencies:
pip install -e .
Train (edit your YAML config or use the default):
python trainer.py --config_path configs/default.yaml
Inference (example):
python inferer.py \ --config_path configs/default.yaml \ --num_samples 2000 \ --model_path mask_class_conditioning_checkpoints/latest \ --num_inference_steps 5
After inference, generated samples are saved as a .pkl in the checkpoint folder noted by --model_path.
Data format & configs
For training, store your dataset in a single .pkl with splits and fields such as
image, mask, class, and optional metadata.
Adjust paths and hyper-parameters in configs/*.yaml.
Reproducibility: set train_args.seed and
train_args.deterministic in config; for inference, optionally pass
--seed.
Resources
Code & checkpoints
MIT-licensed implementation with training, inference, and sample configs.
Citation
@article{yazdani2025flow,
title = {Flow Matching for Medical Image Synthesis: Bridging the Gap Between Speed and Quality},
author = {Yazdani, Milad and Medghalchi, Yasamin and Ashrafian, Pooria and Hacihaliloglu, Ilker and Shahriari, Dena},
journal = {arXiv preprint arXiv:2503.00266},
year = {2025}
}
Contact
For questions about the paper or code, email milad.yazdani@ece.ubc.ca.
Affiliation: University of British Columbia, Vancouver, Canada.
© 2025 The authors. Code: MOTFM (MIT). Datasets referenced: CAMUS, Medical Segmentation Decathlon.
If you use this work in academic projects, please cite the paper. Images belong to their respective copyright holders and are used here for scholarly communication.